Biosynthesis and salvage of queuosine
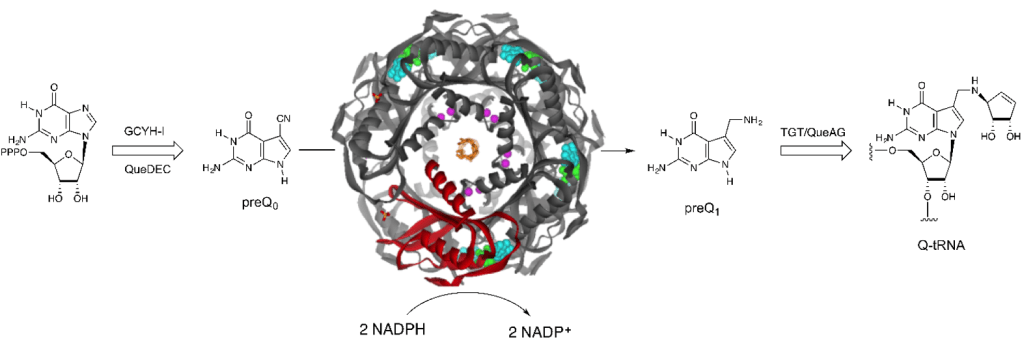
Queuosine (Q) is a 7-deazaguanine modification at the wobble nucleotide in the anticodons of many tRNAs where it ensures fidelity of ribosomal translation by preventing frameshifting. Q is made de novo in bacteria, whereas in human and other mammals it is uptaken from the gut microbiota as a micronutrient. Disruption of Q biosynthesis in some pathogenic bacteria results in loss of pathogenicity due to mistranslation of virulence genes, hence Q biosynthesis enzymes are being pursued as antimicrobial targets. Our lab has been actively engaged in characterizing the Q biosynthesis pathway in bacteria and elucidating the functions, structures, regulation and catalytic mechanisms of the biosynthesis enzymes. Q is important for human physiology and aging, and in particular in the brain where it modulates translation of human genes via modification of human tRNA. Our lab has an ongoing NIH-funded project to study how Q is uptaken and processed in the human body and how it influences brain cell metabolism and function. For publications on this topic, visit here.
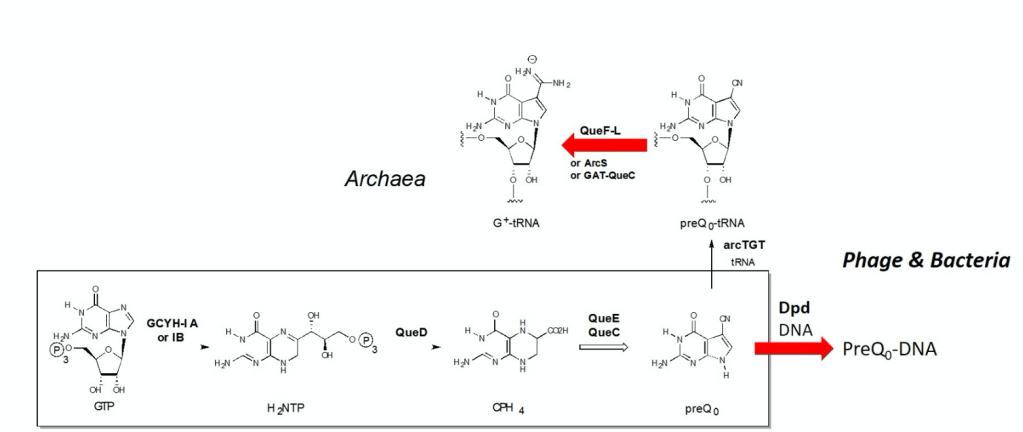
Modification of archaeal tRNA and phage DNA with archaeosine
The positively charged 7-deazaguanosine nucleoside archaeosine (G+) is one of the most highly modified nucleosides found in tRNA. It occurs at position-15 in the dihydrouridine loop of archaeal tRNAs, where it functions in stabilization of the tRNA tertiary structure. G+ and its precursor base 7-cyano-7-deazaguanine are also found in bacteriophage and bacterial DNA where they act as part of a novel restriction-modification defense system. In this project we focus on characterizing the late steps in the G+ biosynthesis pathway in these diverse organisms, and elucidate the catalytic mechanisms and specificities of the diverse enzymes involved. These studies illuminate several exquisite examples of molecular evolution in which diverse chemical transformations emerged in common protein folds, and diverse folds are summoned in nature to facilitate a common biochemical reaction. For publications on this topic visit here.
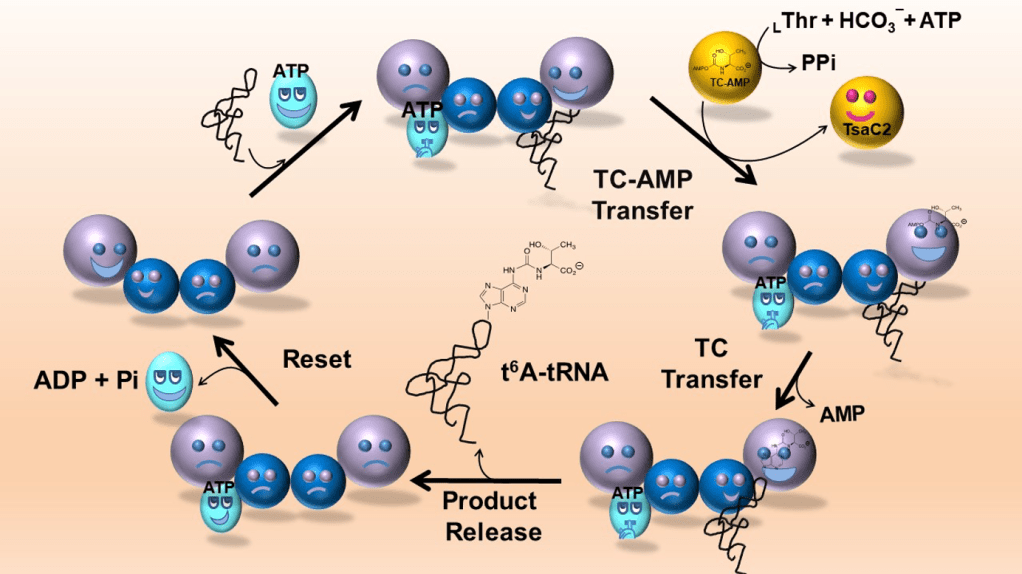
Biosynthesis of N6-threonylcarbamoyl adenosine
In all life forms, position 37 of the anticodon loop of tRNAs decoding ANN codons (N is any nucleotide) is modified with the complex nucleoside threonylcarbamoyladenosine (t6A37). t6A37 is essential for translational fidelity, it promotes correct selection of start codons and prevents aberrant frameshifting during ribosomal translation. In human, defects in t6A37 biosynthesis leads to the renal-neural disease Galloway-Mowat Syndrome. Because the t6A37 biosynthesis enzymes are essential in bacteria, and two of them are unique to the bacterial domain, the t6A37 pathway is a compelling potential target for the development of a new generation of anti-bacterial therapeutics. Our lab has an ongoing NIH-funded project to elucidate the mechanistic and structural basis for t6A37 biosynthesis, and establish the experimental foundation for the development of this critical metabolic pathway as an antibacterial target. For publications on this topic visit here.
Structure-based discovery of new anti-folate antibiotics
This project focuses on developing the prokaryotic-specific GTP cyclohydrolase IB enzyme (GCYH-IB) that catalyzes the first step in queuosine and folate biosynthesis in bacteria, as a new molecular target for antibacterial agents. The bacterial folate biosynthesis pathway harbors several established and clinically validated molecular targets for antimicrobial agents. However, these targets evolved multi-drug resistance. GCYH-IB is found in several human pathogens including Staphylococcus aureus (SA) and Nisseria gonorrhoeae and is an essential enzyme for the survival and growth of SA. GCYH-IB is structurally distinct from the human enzyme that is involved in the biosynthesis of biopterin. These structural differences are exploited in a structure-based approach to discover inhibitors specific for the bacterial enzyme. For publications on this topic visit here.
